Hello, Sign in
My Account
My Account
New customer? Start here
Introduction
Sumoylation is a post-translational modification system that has been the object of scrutiny due to its effect on numerous biological pathways. Analogous to ubiquitin, reversible covalent attachment of SUMO (small ubiquitin-like modifier) to lysine residues in substrate proteins alters the properties of the proteins to which SUMO conjugates. In contrast to ubiquitin, SUMO conjugation does not typically lead to substrate degradation; rather, sumoylation directs multiple effects on many different biological processes, including protein localization and stability, transcriptional activities, nucleocytoplasmic signaling and transport, and genomic replication, as well as the regulation of gene expression and viral reproduction.
More than 100 SUMO-1 target proteins have been reported, and for a host of those, dysregulation is correlated with specific disease states. Protein targets for modification include tumor suppressors p53 (plus regulatory proteins), PML, transcription factors such as receptors for androgen, progesterone and gluticorticoids, c-Jun, c-Myb, and the C/EBP proteins. DNA repair and replication targets include topoisomerase I and II, PCNA and histone deacetylase, with other modified proteins including heat shock proteins and signal transduction molecules.
SUMO Isoforms
In vertebrates, four SUMO isoforms, SUMO-1, SUMO-2, SUMO-3, and SUMO-4, have been identified with distinct function for each. SUMO-2 and SUMO-3 share 95% amino acid sequence identity with each other, but only approximately 47% identity with SUMO-1 (Figure 1). SUMO-4, identified specifically in human kidney, bears homology to SUMO-2/3, suggesting that some SUMO proteins have tissue-dependent functions.

SUMO-1 conjugates to target proteins as a monomer, while SUMO-2 and SUMO-3 conjugate to proteins as higher molecular weight polymers with SUMO-1 terminating further SUMO addition. The in vivo functions of SUMO-2/3 modifications appear to segregate from those of SUMO-1. All SUMO isoforms are attached to substrate proteins through a biochemical pathway similar to that of ubiquitination.
The Molecular Role of SUMO Modification
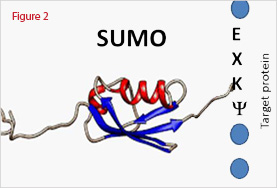
For SUMO modification, a consensus sequence Ψ-K-x-D/E (where lysine is the modified amino acid, Ψ is a hydrophobic residue and x is any amino acid residue) is required for SUMO modification (Figure 2 and Table 1).
| Protein name | N | Core | C | Ubc9 Binding | Molecular module |
|---|---|---|---|---|---|
| RanGAP1 | HMGL | LKSE | DKV | Yes | Nuclear import |
| PML (1) | HQWF | LKHE | ARP | Yes | Tumor Suppression |
| PML (2) | PRKV | IKME | SEE | Yes | Tumor Suppression |
| Sp100 | RLVD | IKKE | KPP | ? | Chromatin remodeling |
| p53 | KKLM | FKTE | GPD | Yes | Tumor Suppression |
| lkB alpha | PRDG | LKKE | RLL | Yes | Signal transduction |
| c-Jun | RLQA | LKKE | PQT | Yes | Transcription activation |
| IE2 (1) | MLPL | IKQE | DIK | Yes | Viral regulation (CMV) |
| IE2 (2) | KQED | IKPE | Yes | Viral regulation (CMV) | |
| HSF2 | DSGI | VKQE | RDG | Yes | Transcription activation |
| AR | PHAR | IKLE | NPL | Yes | Transcription activation |
The mechanism involved in maturation and transfer of SUMO to target substrates comprises four steps: maturation, activation, conjugation, and ligation. First, the SUMO protein is cleaved by SUMO-specific carboxyl-terminal hydrolase to produce a carboxyl-terminal diglycine. SUMO modification then proceeds by a three-step enzyme shuttle, a process composed of an E1-E2-E3 enzyme catalyzed cascade of activation (E1), conjugation to E2 (UBC9), and finally addition to the terminal amino group of a lysine side chain in target proteins, which may require E3 (ligase). E1 and E2 appear to be conserved for the SUMO proteins, while several different SUMO ligases (E3) have been identified in higher eukaryotes. Despite the sequence differences, the activation enzyme E1 and conjugation enzyme E2 do not discriminate among SUMO molecules. SUMO covalent linkage to the substrate protein is labile, and SUMO-deconjugation enzymes add another level of regulation to the sumoylation process.
Biological Functions of SUMO modification
The number of proteins and pathways affected by SUMO regulation, and the function thereof are extensive. Herein are featured just a few key examples in the areas of protein localization, genetic regulation, transcriptional activities, and viral reproduction.
Subcellular Localization
Sumoylation participates in regulating the subcellular partitioning of dozens of substrate proteins. As an example, RanGAP1, a cytoplasmic protein facilitating protein transport across the nuclear pore complex, is localized to the nuclear pore complex upon SUMO modification. The SUMO E2 conjugation enzyme and the E3 ligase are enriched at the nuclear pore complex. Nuclear bodies containing multiprotein complexes have yielded deep insight into the role of SUMO modification in protein recruitment and localization. Sumoylation has been found to be required for the subcellular localization of many proteins found in PML nuclear bodies, while the sumoylation state of other proteins does not control their localization to the PML nuclear bodies, but does affect their ability to recruit binding partners. For example, the sumoylated forms of PML and Sp100 are found only in the nucleus; this modification is mandatory for PML localization. On the other hand, the targeting and accumulation of Sp100 in these bodies does not depend upon sumoylation as revealed through studies mutating the target lysine.
SUMO and the Chromosome
Partitioning of chromosomes into replicated cells is an intricate process dependent upon the precise scheduling of sister chromatid assembly and separation. Observation of dysregulation of this process in yeast SUMO-1 mutants, resulting in defective mitosis and chromosomal segregation, was one of the first findings describing the role of SUMO modification in maintaining genetic integrity. Sumoylation has also been implicated in the recruitment and retention of proteins to the kinetochore, the protein complex that forms at the centromere and gathers microtubules for anaphase separation.
Sumoylation of tumor suppressor and repair proteins points to the role of SUMO-1 in preserving genomic stability. p53 and Mdm2 are both sumoylated, with functional effects on activity. Proteins in the Wnt signaling cascade (e.g. axin, β-catenin, LEF/Tcf-4) are also targets of sumoylation. A number of DNA repair enzymes are also subject to regulation by sumoylation. Sumoylation is also been implicated in the repair of DNA damage mediated by topoisomerase II.
SUMO and Transcription Regulation
Sumoylation of proteins involved in the transcription machinery is involved in regulation of gene expression, primarily through repression, with the noticeable exception of heat shock factors, which are upregulated. Transcription factors inhibited by SUMO modification of the substrate lysine include STAT1, c-Jun, CEBP alpha, c-Myb, IRF-1, SREBPs, SRF, Elk, AP1, AP2, androgen receptor (AR), glucocorticoid receptor (GR), and progesterone receptor (PR), and huntingtin.
Many transcription factors are regulated by association with PML nuclear bodies, and assembly of these structures requires sumoylation of the PML protein. The PML gene, involved in the chromosomal translocation of acute promyelocytic leukemia, encodes a protein which localizes to PML Oncogenic Domains (PODs), subnuclear macromolecular structures. The effect of sumoylation upon transcriptional activity is therefore broadcast widely downstream. Sumoylation of PML recruits corepressor Daxx to PML nuclear bodies, thereby removing Daxx-mediated repression of these genes. Similarly, sumoylation of PML directs p53 to PML nuclear bodies, which stimulates transcriptional activity of p53; given the ubiquitous role of p53 in cellular signaling pathways, the ramifications of p53 sumoylation are significant.
The effects of SUMO on transcriptional activity may also operate indirectly – discovery of sumoylation of a number of transcription co-activators and co-repressors, such as GRIP1, SRC-1, and histone deacetylases (HDACs) suggests the complexity of sumoylation in transcriptional network regulation.
SUMO and Viral Infections
The modification of specific viral proteins by SUMO has been demonstrated to play a role in viral reproduction. Sumoylation of major-immediate-early proteins for human cytomegalovirus, herpes virus, Epstein-Barr, adenovirus, and papillomavirus have been shown to enhance transactivation of key proteins in viral reproduction, and to effect localization of the viral proteins to the host nucleus. Sumoylation of host proteins may provide a cellular environment promoting or deterring viral infection. Work in this area is accelerating, given the potential to understand and interrupt by novel means viral infections via targeting of the sumoylation pathway.
Conclusion
The array of biological functions in which sumoylation is involved place this modification among those such as phosphorylation that have far-reaching effects on cellular function. The continuing interest in this small protein with big effects on cellular networks bound up with health and disease augurs for new discoveries and novel therapeutic approaches to diseases impacted by SUMO protein substrates.