Hello, Sign in
My Account
My Account
New customer? Start here
In 2004, Avram Hershko, Aron Ciechanover and Irwin Rose won the Nobel Prize for Chemistry for their contribution to the understanding of the ubiquitin proteasome role in cell biology. The ubiquitin pathway is the principal mechanism for protein catabolism in the mammalian cytosol and nucleus. The tightly regulated pathway affects a wide variety of cellular processes, and defects in the system can result in multiple important human diseases. The ubiquitin pathway is central to:
Pathologies that result directly or indirectly from aberrations in the ubiquitin pathway are:
Degradation of a protein via the ubiquitin pathway involves two successive steps: tagging of the substrate protein by the covalent attachment of multiple ubiquitin molecules (conjugation); and the subsequent degradation of the tagged protein by the 26S proteasome. Therefore, the primary functions of ubiquitin have been associated first with protein housekeeping and turnover as well as antigenic-peptide generation. A significant regulative function of protein modification by ubiquitin has been unveiled in DNA repair and endocytosis. These functions are controlled by the number of ubiquitin units attached to proteins (mono- or poly-ubiquitination) and the type of ubiquitin chain linkage. The reversible linkage of one molecule of ubiquitin or ubiquitin-like molecules such as SUMO and NEDD8 to proteins is involved in critical biological processes, including protein localization, transcriptional activity, nucleocytoplasmic transport and genomic replication.
The Ubiquitin Conjugation Mechanism
Ubiquitin, an essential protein of 76 amino acids, represents one of the most conserved polypeptides in eukaryotes, and only four of its amino acids differ among yeast, plants and animals. Ubiquitin is expressed as three different precursors: a polymeric head-to-tail concatemer of identical units (polyubiquitin); and two N-terminal ubiquitin moieties, UbL40 and UbS27, that are fused to the ribosomal polypeptides L40 and S27, respectively. Specific endopeptidases cleave these precursor molecules to release ubiquitin moieties that are identical in sequence. In budding yeast, the ribosomal fusion proteins are responsible for the bulk of the free ubiquitin pool. Before its proteolytic separation from L40 and S27, the ubiquitin moiety acts as a chaperone and thus facilitates ribosome assembly. Only a few amino acids in ubiquitin are vital for its function, however it seems that the highly conserved ubiquitin gene sequence is driven not only by functional properties of the ubiquitin protein, but also by the propensity of the polyubiquitin locus to act as donor or substrate for recombination to maintain the non-allelic ubiquitin genes.
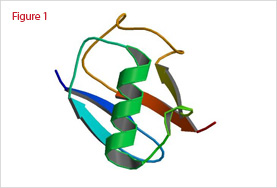
The 3D-structure of ubiquitin solved by X-ray diffraction and NMR has shown that its structure is highly compact and rigid (Figure 1). Only the last four C-terminal amino acids are protuberant and unstructured. Within these four residues, Bly-76 is activated in an ATP-dependent manner for subsequent ubiquitin ligation with the epsilon amino group to form an additional peptide bone to generate multi-ubiquitin chains.
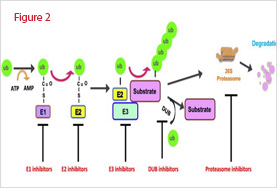
The ubiquitin conjugation cascade starts with the activation of the carboxyl group of Gly-76 of ubiquitin by the ubiquitin-activating enzyme E1 (UBA1), the only E1 enzyme coded by the yeast genome. This ATP-dependent reaction generates a ubiquitinyl adenylate intermediate bound to the E1 enzyme. The activated ubiquitin is then transferred to a thiol group of an active site Cys of a ubiquitin-conjugation enzyme E2. Eleven E2 enzymes (Ubc1-8, 10, 11, 13) have been identified in the yeast genome and many more in higher organisms. They all share an active-site ubiquitin-binding Cys residue and are distinguished by the presence of a UBC domain required for binding of distinct E3 enzymes. Finally, E2 enzymes transfer the ubiquitin moiety, either directly to a protein substrate’s internal Lys residue. The first case is catalyzed by E3 ligases encompassing a RING finger domain, which serve as scaffolds to bring together E2 enzymes and substrates. The second case is catalyzed by HECT domain E3 ligases. Each E2 enzyme interacts with numerous E3 ligases, thus expanding the number of substrates targeted. Once the protein substrate is mono-ubiquitinated, a polyubiquitin chain is formed through the same ubiquitination conjugation cascade (Figure 2).
The 26S Proteasome: a Multi-subunit Complex for Protein Degradation
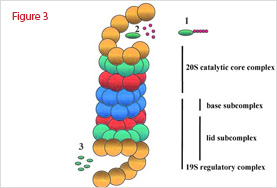
The 26S proteasome is a large 25,000-kDa complex made of two copies of at least 32 different subunits. The overall structure can be divided into two major subcomplexes: the 20S CP that contains the protease subunits, and the 19S RP regulatory particle that regulates the function of the 20S CP (Figure 3). The structure of the 20S proteasome is well conserved in all organisms. The barrel-shaped complex consists of four stacked heptameric rings that form a channel in which the unfolded substrate goes through. Each ring is composed of seven distinct subunits. The alpha-subunits form the two outer rings, which have a regulatory role with the 19S particle. The beta-subunits form the two inner rings, which contain the proteolytic active sites within a sequestered chamber. The 19S RP particle is made of two subunits, a lid and a base. While the lid recognizes ubiquitinated substrates, the base unfolds protein substrates and threads them into the catalytic chamber of the 20S particle. The base contains siz ATPases that cap the 20S particle by interacting with the alpha-subunits. Altogether with ancillary proteins, the 19S complex regulates the opening of a gated channel into the 20S core to free access to the proteolytic chamber otherwise inactive in the 20S particle alone.
Recycling Ubiquitin and Deubiquitination Enzymes
Ubiquitin moieties attached to a protein substrate degraded by the 20S complex are released from the proteasome and recycled back into the ubiquitin pathway. The deubiquitination activity specifically associated with the proteasome complex has not been yet identified but could be attributed to p37, a protein in Drosophila. Deubiquitinating enzymes (DUBs) serve to edit the ubiquitin chains and remove ubiquitin from protein substrates before degradation. More than 17 DUB enzymes have been identified in eukaryotes. Based on their molecular size, sequence homology and active site residues, DUBs are categorized as UCHs (ubiquitin COOH-terminal hydrolyses) or UBPs (ubiquitin-specific proteases). UCHs are generally small enzymes (20-30-kDa) that remove short and flexible peptide chains from the COOH terminus of ubiquitin. UBPs on the other hand belong to a more diverse group of enzymes with a larger molecular mass (100 kDa). UBPs cleave the isopeptide bond linking Ub-Ub or Ub-protein and also biosynthetic linear fusions of ubiquitin. DUBs have several roles in maintaining the steady-state levels of free ubiquitin and in affecting the stability of ubiquitin-conjugated proteins. DUBs generate and recycle ubiquitin, edit polyubiquitin chains, and aid in the proteasome-dependent degradation of proteins.
Concluding remarks
The ubiquitin conjugation mechanism includes about 50 E1 and E2 enzymes, 500 to 700 E3 enzymes, and 70 to 100 DUB proteins. From our current knowledge of the protein degradation field, numerous ligases have emerged as tempting anticancer targets. Subsequent target validation and drug screening have been initiated by pharmaceutical companies. Although preliminary studies have shown exciting results on cancer cell models, there are still many unknown effects of the potential drug candidates to investigate. With each ligase degrading multiple proteins, it is difficult to predict every secondary effect a drug may trigger by inhibiting only one ligase. In this aspect, E3 enzymes may be better targets than E1 or E2 enzymes, which are likely less specific than E3s due to their lower number. DUB enzymes are considered as the next set of enzymes to target in the ubiquitin pathway. Another family of enzymes of high interest for drug development is the ubiquitin-like molecules. Numerous substrates for SUMO and NEDD8 have been identified by proteomics approaches as druggable targets. Nonetheless, it will take a serious commitment to unveil the detailed mechanism of ubiquitin-mediated protein modification and to understand the therapeutic ramifications of pharmaceutical intervention.