Hello, Sign in
My Account
My Account
New customer? Start here
Antibody Production: Starting Out Right
Antibodies are powerful tools to profile protein function, interaction, cellular location, expression, and post-translational modification, and as such are indispensable to modern biological research. The first step in antibody production consists in making the decision regarding the type of immunogen: protein or peptide. The ability to produce an immune response in an animal to a peptide conjugated to an appropriate carrier protein circumvents in many cases the time and expense associated with isolating or expressing and purifying native protein. Additionally, the use of a peptide permits one to produce epitope-specific polyclonal antibodies, an extra measure of selectivity not possible when immunizing with whole protein. The substantial reduction in cost and time are compelling incentives that have fueled the rise of anti-peptide antibodies in both basic and applied research. That said, there are a number of conditions that go into designing the peptide antigen and producing the antibody. Understanding those issues while working with the antibody production team enhances the likelihood of success. The Epinetix™ epitope design procedure has been developed by ADMEbio scientists, compiling the expertise of different scientific fields within the same team. This multi-pronged approach is essential to ADMEbio antibody production, and is now the subject of this review.
Defining The Right Expectations For The Project
Whether the antigen is a peptide or protein, the end user must choose whether a polyclonal or monoclonal antibody best matches their specific applications and experimental requirements.
Polyclonal antibodies will recognize multiple epitopes of an antigen. Since the minimum length of an epitope is thought to be 4-6 amino acids, even a peptide may contain several epitopes. Polyclonal antibodies are more likely to maintain recognition of antigenic epitopes even when modest changes in conformation or aggregation occur. Polyclonal antibodies are also capable of recognizing different epitopes with different affinities. Because of this ability, polyclonal antibodies have a broader range of potential applications than monoclonal antibodies.
Monoclonal antibodies recognize only one epitope of the antigen and are highly specific to that particular antigen. They generally yield much lower background because of this specificity. Monoclonal antibodies are also highly homogenous and allow for unlimited production with sustained specificity.
Whether producing a polyclonal or monoclonal antibody, much effort goes into selecting a peptide with high antigenicity, meaning that the particular amino acid composition and sequence strongly stimulate the immune system to produce antibodies that will bind to the peptide. This is a crucial first step for the project, and the step over which the most control can be exerted. Greater detail will follow regarding specific methods for this step. However, it is important to understand that with the skillful and experienced application of design principles to peptide selection, an anti-peptide antibody that recognizes the native protein may be obtained for approximately 75% of projects. The failures may be assigned to a number of reasons. The cognate peptide sequence may not be readily accessible to the antibodies, because it is buried in the protein structure. The target sequence within the protein may be anchored within secondary structural elements that render it highly inflexible. A flexible free peptide may generate antibodies to structures that do not occur within the protein. It is also possible that anti-peptide antibodies will not bind to certain denatured forms that render the target sequence inaccessible. Finally, it is important to remember that peptides represent linear B-cell epitopes. Vaccine research combined with three-dimensional structural studies has shown that many epitopes are discontinuous, i.e. conformational in nature, defined by the proximity in space of amino acids that are far apart in primary sequence. This library of epitopes is inaccessible to the anti-peptide antibody approach.
Biophysical Parameters For Epitope Prediction
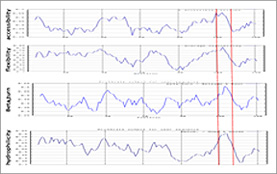
The selection of candidate peptides for antibody production from the full-length protein sequence has been subjected to a dizzying array of predictive algorithms, all of which work from different facets of an underlying principle: The immune system produces antibodies against "features" that are present on the "surface" of the protein. Amino acids can be ranked, based on previous structure knowledge, according to their respective probabilities in promoting attractive structural features for antibody recognition (i.e. proline’s role in promoting B-turns) and in promoting surface exposure (i.e. the tendency of highly hydrophilic residues such as arginine and lysine to position between protein and the aqueous environment). 1
Generally, the average values for these physiochemical parameters are plotted over a window of 6-8 amino acids moving along the protein sequence. Regions that show a consensus among the different parameters are then further evaluated against other criteria for final selection of the peptide [Figure 1].
Antibody Production: The Epinetix™ Epitope Prediction Procedure
With the Epinetix™ epitope prediction procedure, ADMEbio provides customers with a report that visually summarizes the biophysical information, and makes accompanying recommendations about the best candidates for antibody production. Such criteria include the following:
1. Length
The length of the peptide should be between 12 and 25 amino acids according to ADMEbio experience. Longer peptides may adopt non-native structures that give rise to antibodies that do not recognize the native protein. Shorter peptides do not provide sufficient epitopic diversity, thereby producing a constricted range of antibodies, limiting the opportunity to generate antibodies that recognize the native protein. In the case of extremely short peptides selected from the interior of the protein sequence, for example, the overwhelming majority of the antibodies produced may be against the charged N or C terminal residue carrying the respective amide or carboxyl functional group, which would impair recognition by antibodies of the sequence in a native protein where the residue is linked to its flanking neighbors by a peptide bond. The exception to this length requirement occurs in the case of antibodies generated to phosphor or other modification-specific sites. Phosphorylation of a single residue produces minor conformational changes in an epitope. It is desirable to “focus” antibodies on this site, so peptides of 8-12 amino acids with the modification centrally located are selected to enhance the production of phosphospecific antibodies. ADMEbio is experienced in special synthesis and purification protocols for antibodies specific to a range of post-translational modifications, including phosphorylation, methylation and acetylation.
2. Post-translational modifications and topology
Unless one is interested in generating antibodies that are modification specific, peptides candidates must be carefully screened to avoid those incorporating known modification sites, since the modification may block recognition by the antibodies of the native site if it is modified. Cysteine disulfide bonds can introduce unique structural features, bulky glycosylation sterically hinders access by antibodies, and a host of more than 200 known protein modifications may introduce unknown perturbations in the structure of either the peptide or the protein that may impair the transfer of peptide recognition to protein recognition by an anti-peptide antibody. Online tools for screening peptides for these modifications are available. 2
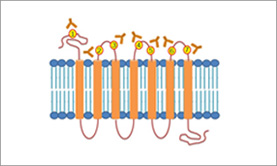
When designing peptides for antibodies against transmembrane proteins such as G-protein coupled receptors, it is absolutely critical to avoid those regions that span the membrane, since the antibody does not have the ability to access them in the native protein. The membrane spanning regions may have been described in publications; alternatively, online tools are available to assist in defining these regions [Figure 2]. 3 Whether to choose a peptide from the extracellular or cytoplasmic loops will depend upon the applications and the objectives defined by the researcher.
3. Conjugation
Peptides less than 25 amino acids in length are usually conjugated to keyhole limpet hemocyanin (KLH) or some other immunogenic large molecular weight protein. Conjugation is typically done through a single cysteine present at the N or C terminus of the peptide. Internal cysteines are not desirable conjugation sites because the full peptide containing all epitopes will be partially occluded by the carrier protein and therefore will restrict generation of a full complement of antibodies. Given these requirements, three scenarios are possible:
4. Ease of peptide synthesis and solubility
5. Sequence Homology
To enhance specificity, peptide sequences must be tested for homology against other proteins from the target species. This assures that peptides holding epitopes in common with proteins other than the target protein are removed from consideration. Other cases where homology may come into play include a requirement that an antibody work in more than one species, or that an antibody be isoform specific. Candidate peptide sequences can be blasted against either the NCBI or Swiss-Prot protein databanks, and the results are then easily surveyed to determine which candidate best meets these criteria.
6. Structural Information
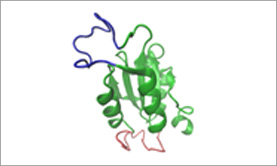
It is often the case that a structure for a target protein is known from X-ray crystallography or NMR study, and the coordinates are available from a public databank such as PDB. There are also a number of free prediction programs online that predict structures from submitted sequences based on homology modeling. 4 Viewing 3D structures with color-coded epitopes can help in either identifying or confirming that peptide sequences are positioned on the surface of the protein, where they have the highest likelihood of producing an antibody [Figure 3].
At Last: Peptide Purity And Quantity
The quantity of peptide required will depend upon the immunization and boosting schedule, and the number of animals to be immunized. While as little as 0.1 mg per animal has been used in special protocols, it is generally recommended to plan for 1-2 mg of peptide per animal. If peptide affinity purification of the antibody is required, an additional 5 mg of peptide should be allotted.
Peptide purity >70% is generally sufficient for production of good anti-peptide antibodies. Peptides that were truncated during synthesis are included in these impurities, and contain virtually the same epitopes as the parent peptide. While HPLC purification of peptides to >90% or >95% may add 5-20% to the cost of the project depending upon the peptide, such stringency may be called for where the antibody must discriminate between proteins differing in only a few amino acids within the epitope. Production of reagent antibodies has historically employed purification against a protein G or protein A matrix. However, it is increasingly common to affinity purify the antibody against the immunizing peptide itself, coupled to a resin. This tends to reduce the number of contaminant antibodies that would otherwise co-elute, improving the background signal. Peptide affinity purification costs somewhat more and reduces the yield of antibody (IgG fraction) up to 95%.
Some researchers seek to reduce project costs by co-immunizing animals with multiple peptides. This two-for-one approach is intended to enrich the diversity of antibodies that are produced and to short-circuit potential failure if a single antigen does not produce an immune response. In some cases a good polyclonal antibody that can capture the protein from multiple epitopes is produced. However, in practice not all antigens are equal in immunizing potential. The immunodominant peptide may produce the vast majority of antibodies, and in cases like this little ground is gained for the added expense. Consultation with the peptide and antibody production teams prior to choosing a strategy will help in making the best decision regarding this option.
In Closing
This article has reviewed key aspects of peptide design for the production of antibodies. With so many aspects to consider and literally thousands of peptide sequences available from a full-length protein, epitope prediction programs have come to the fore as a powerful means to winnow down the choices to a handful for consideration. Although such automated computational methods are becoming indispensable to high-throughput production of commercial scale antibody collections, the field still demands the art of a practiced team to consistently produce successful outcomes. For custom antibody production, the need for a strong collaboration and effective communication with the vendor remains as essential as ever. The ADMEbio Epinetix™ epitope design procedure employs the expertise of a multi-scientific background production team to generate the best reagents for our customers.
References